Habitat and Connectivity
Habitat.RmdIntroduction
In this vignette, we explore the database linking OCS-GE (French GIS layer for Occupation du Sol à Grande Échelle) labels with species labels.
We will analyze how different land cover types act as habitats with varying qualities or resistances for different species, starting with the Wood mouse (Apodemus sylvaticus).
Defining Habitat
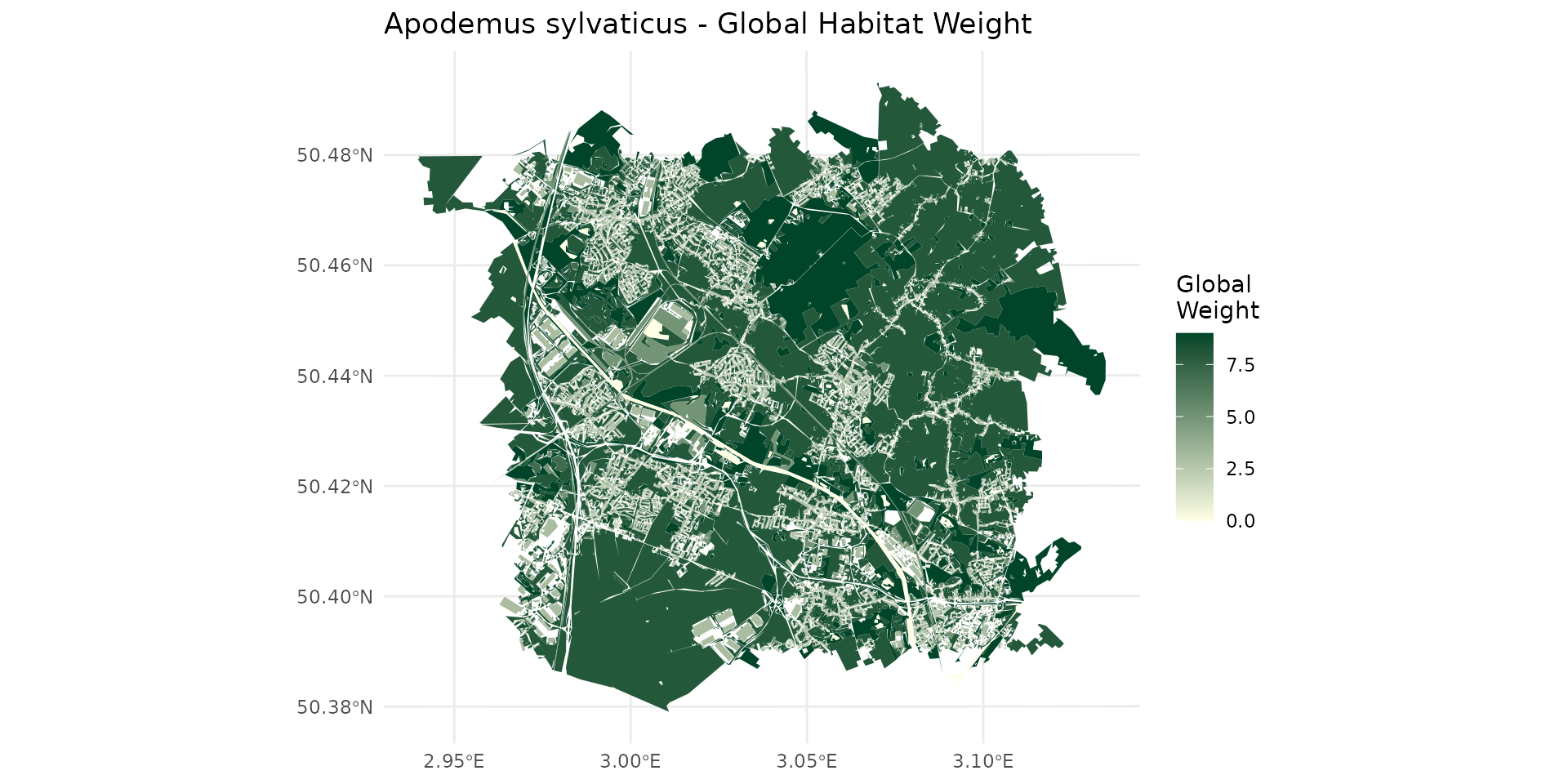
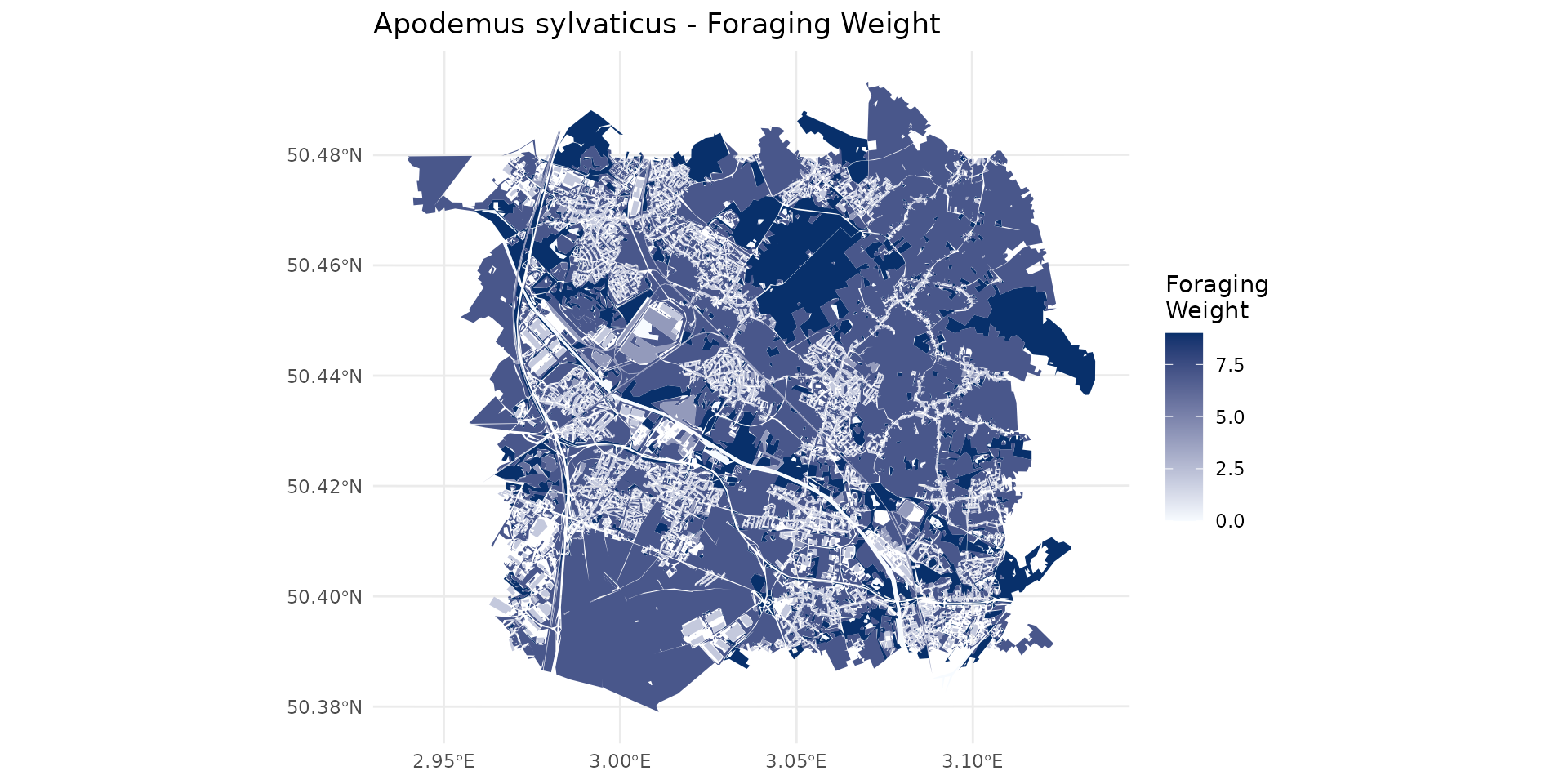
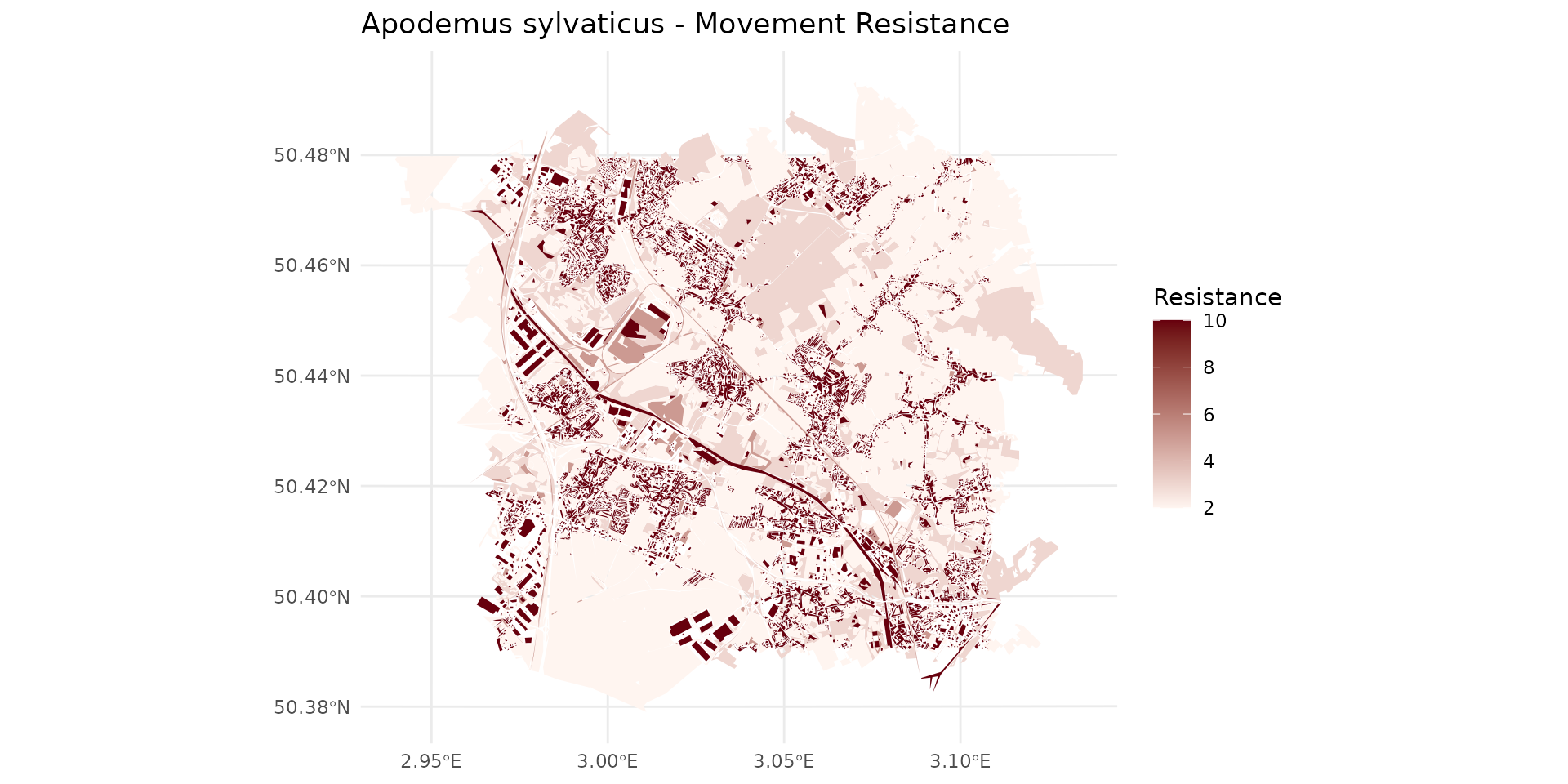
Weighted species habitat with OCS-GE layer for Apodemus sylvaticus
First, we load the necessary datasets provided by spacemodR:
- The species dictionary assigning weights to land cover codes.
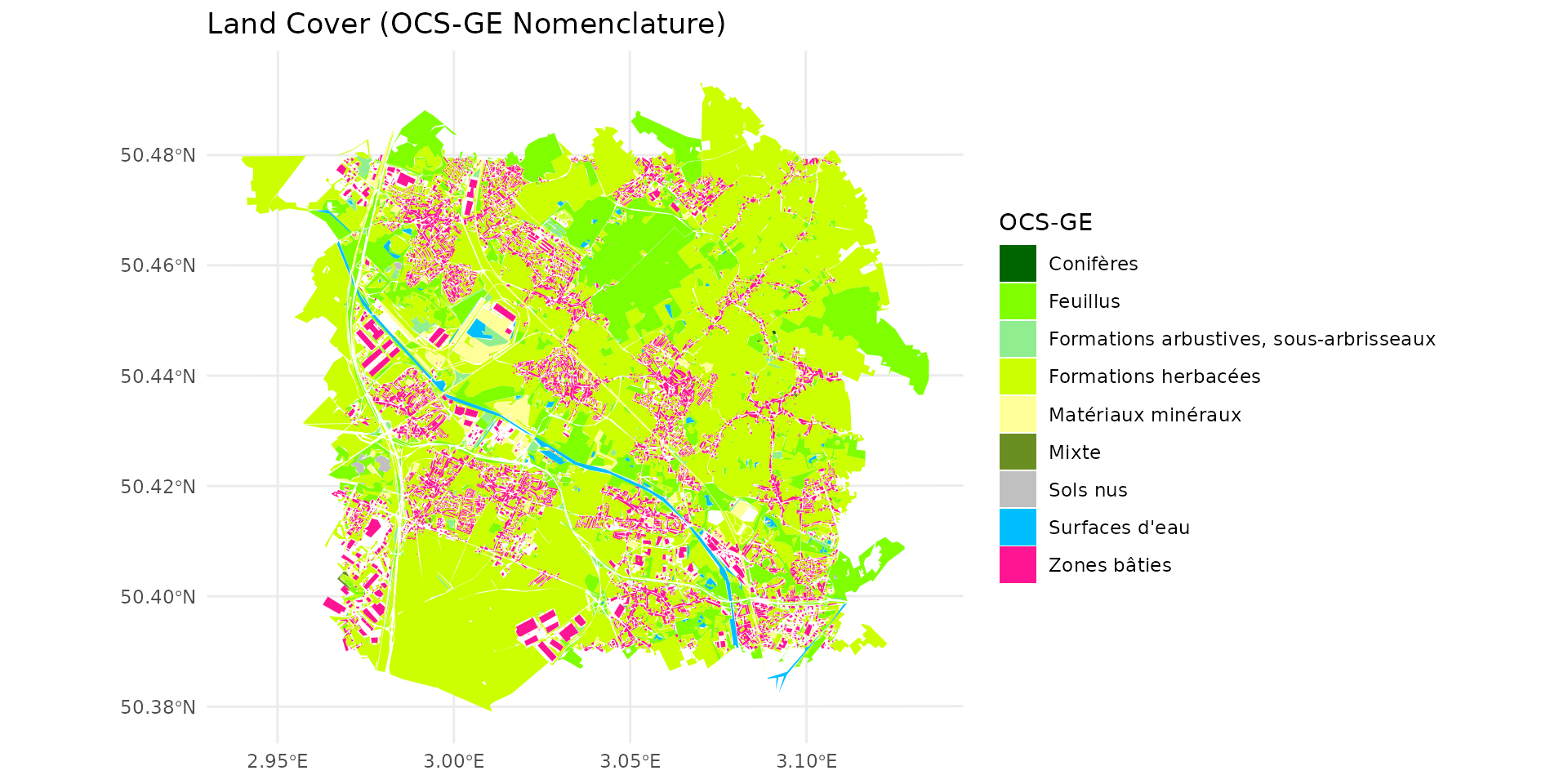
- The OCS-GE nomenclature reference (containing names and specific hex colors).
- The spatial layer for OCS-GE (here, we use the ocsge_metaleurop dataset as our base map).
# Load the species dict and reference tables
data("ocsge_species_dict")
data("ref_ocsge")
# Load spatial data
data("ocsge_metaleurop")
sf_ocsge <- ocsge_metaleuropWe filter our species dictionary for Apodemus sylvaticus and join it with the reference nomenclature to link the OCS-GE codes to their descriptive labels and predefined colors.
# Filter for Apodemus sylvaticus
dfhab_Apsy <- ocsge_species_dict[
grepl("Apodemus_sylvaticus",ocsge_species_dict$nom_espece, ignore.case = TRUE),
]
# Join ref_ocsge with the species dictionary.
# ref_ocsge$code_cs_ (with spaces) matches ocsge_species_dict$code_cs
df_merged_Apsy <- ref_ocsge %>%
dplyr::left_join(dfhab_Apsy, by = c("code_cs_" = "code_cs"))
# Join with the spatial sf object. Assuming sf_ocsge uses 'code_cs' (without spaces)
sf_Apsy <- sf_ocsge %>%
dplyr::left_join(df_merged_Apsy, by = "code_cs")Understanding Habitat Values
By examining the weight_global (wg), we can classify the landscape into different habitat qualities for the species:
- Non-habitat area (wg = 0):
df_merged_Apsy %>%
filter(weight_global == 0) %>%
pull(nomenclature) %>%
unique()
#> [1] "Surfaces d'eau" "Névés et glaciers"- Very Poor habitat (0 < wg <= 3):
df_merged_Apsy %>%
filter(weight_global > 0, weight_global <= 3) %>%
pull(nomenclature) %>%
unique()
#> [1] "Zones bâties" "Zones non bâties"
#> [3] "Autres formations non ligneuses"- Poor habitat (3 < wg <= 7):
df_merged_Apsy %>%
filter(weight_global > 7) %>%
pull(nomenclature) %>%
unique()
#> [1] "Feuillus"
#> [2] "Conifères"
#> [3] "Mixte"
#> [4] "Formations arbustives, sous-arbrisseaux"
#> [5] "Formations herbacées"- Good habitat (wg > 7):